Peter Jones, PhD, and Charis Himeda, PhD, found themselves unexpectedly in the media spotlight last week after publication of their paper on the application of CRISPR technology in FSHD cells. In response to the public’s interest in the story, they prepared the following “primer” to explain basic concepts and clarify some points about which people might get confused.
Cell biology
In the simplest, nutshell version, the genome consists of all the DNA in a cell, which exists in the form of chromosomes. Genes are sequences of DNA that code for RNAs, which, in turn, code for proteins. When RNA and protein are being actively produced from a gene, the gene is referred to as expressed (“on” or “active”). Proteins, in general, are the machinery that run a cell. Some of this machinery is common to all cell types (things that every cell needs) and some of it is specialized for specific cell types (certain proteins allow a cell to function as a muscle cell as opposed to a blood cell or a liver cell).
Background on FSHD genetics
About half the human genome consists of repetitive DNA sequences that are normally repressed (collectively referred to as the “repeat genome”). The FSHD disease locus is within the repeat genome, at a large repeat called D4Z4. D4Z4 repeats are present in hundreds of copies at several different chromosomes. FSHD is caused by a shortening of the D4Z4 repeat (to fewer than 10 copies) on a specific type of chromosome 4. This shortening causes a loss of repression in the region, which allows the DUX4 gene (harbored within each copy of D4Z4 and normally silent in somatic cells) to be turned on from a single pathogenic repeat unit (the last unit of the array). The abnormal expression of DUX4 in skeletal muscle leads to FSHD.
Background on CRISPR/Cas9
CRISPR/Cas9 is a gene-editing technology that involves two key components:
- the Cas9 protein from bacteria and
- a guide RNA that targets Cas9 to a specific sequence within the genome.
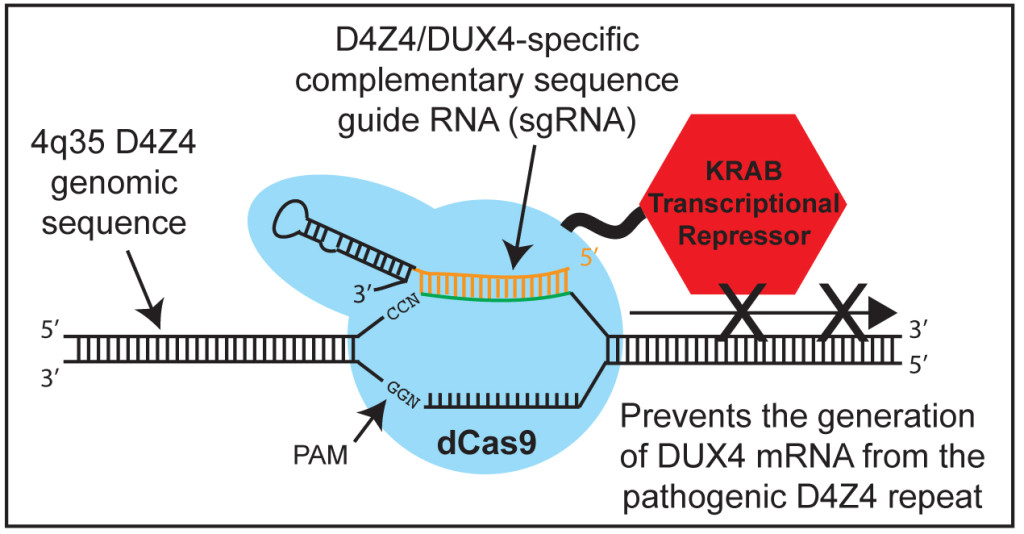
Normal Cas9 (the version most commonly used) is an enzyme that cuts DNA, allowing subsequent insertion of a changed or new DNA sequence into the genome at a specific location (thus, the phrase “gene editing technology” associated with the typical CRISPR/Cas9 approach). We used a non-cutting version of Cas9 (“dead Cas9” or dCas9) fused to another protein (KRAB) whose normal function is to turn off gene expression. In principle, dCas9-KRAB enables one to target and specifically silence the expression of any target gene.
CRISPR in the context of FSHD vs other diseases
Most diseases are caused by mutation(s) in a single protein-encoding gene, which results in the production of a dysfunctional protein or no protein at all. Thus, researchers studying these “typical” diseases are using Cas9 to cut out the mutated part of the disease gene in order to replace it with the correct DNA sequence. For FSHD, we don’t need to replace a missing/dysfunctional protein; rather, we need to stop production of a toxic protein. Hence, CRISPR inhibition with dCas9-KRAB (as opposed to editing with Cas9) may be uniquely suited to FSHD.
Major findings from our study
Most genes are present in only two copies (one from each parent) and are very similar; therefore, targeting these sequences is conceptually quite simple. By contrast, only one specific copy of D4Z4—the last repeat unit on a specific type of chromosome 4—expresses DUX4, causing FSHD. This introduces a degree of complexity for genome targeting that is not normally encountered. It was unclear whether the existing CRISPR technology could be used to target a single pathogenic copy of D4Z4, amongst hundreds of other nearly identical sequences in the genome. In this study we showed that we could, in fact, target a repetitive region of the genome and repress expression of the pathogenic DUX4 gene using dCas9-KRAB.
Typically, researchers use Cas9 to edit problematic DNA sequences, and that approach is also being attempted for FSHD. However, cutting a repeat sequence that is present in hundreds of nearly identical copies could be very dangerous or even catastrophic for a cell. A key aspect of our study is that all D4Z4 repeats are normally in the OFF (unexpressed) state, and only in FSHD is the pathogenic D4Z4 in an ON (expressed) state, leading to abnormal expression of the DUX4 gene. We therefore decided to use dCas9-KRAB to try to return all D4Z4s to the OFF state. Those already OFF would remain OFF, and those ON would be switched to OFF, including the one D4Z4 that leads to FSHD. Our study shows that we can design and utilize guide RNAs to target the dCas9-KRAB repressor to the FSHD disease locus, effectively returning the DNA to its OFF state and repressing expression of the pathogenic DUX4 gene.
Our study is:
- The first to show that the repeat genome can be effectively targeted using CRISPR technology;
- The first use of CRISPR inhibition for any human disease (as opposed to CRISPR editing, which is being widely used in other disease studies by other labs). Note: we did not develop the CRISPR inhibition tool (dCas9-KRAB)—it was developed by other labs;
- The first use of CRISPR technology in primary human muscle cells (which are closer to the cells in a patient’s body than the more commonly used “immortalized” cell lines).
What makes this finding different from other work done to prevent/treat FSHD?
Many researchers are investigating ways to silence the expression of DUX4 or its downstream pathways that lead to FSHD. By contrast, we aimed to overcome the underlying genetic defect by returning the DNA at the FSHD disease locus to its normal repressed state. This approach has several advantages:
- It should correct aspects of the disease caused by loss of repression at the disease locus, including the pathological expression of DUX4;
- It should correct all aspects of the disease that are caused by DUX4 expression; and
- Repressive changes in the structure of the disease locus have the potential to be passed on to subsequent generations of cells.
Specificity of the CRISPR technology is also an issue. In addition to binding their intended target sequence, guide RNAs also have an affinity for similar sequences in the genome, and can recruit the Cas9 cutting enzyme to these off-target locations. By using dCas9, we avoid the serious concern of cutting the genome at unintended places.
What do you hope will come of this? Is this a step to preventing FSHD entirely?
We believe that this work has laid the foundations for using CRISPR technology to treat FSHD in the future. Rapid advances in genome-editing platforms are actively underway, and will hopefully lead to more effective and specific correction of many diseases. For example, there has already been successful development of new Cas9 enzymes that are amenable to viral delivery, which is critical for moving towards therapy.
Our work has demonstrated that, in principle, such a strategy is feasible for FSHD, by showing that the CRISPR system can be used to target and correct a very unusual part of the human genome. Ultimately, more work needs to be done before CRISPR technology for FSHD or any other disease will be ready to move into a clinical setting. It is imperative for us to have a better understanding of off-target effects and the long-term implications of gene editing/modification. Luckily, progress in the treatment of any disease is progress for all, since many therapies and technologies are broadly applicable, as are lessons learned at all stages of therapy development.
Putting CRISPR in the context of current research funding
The CRISPR system was discovered in bacteria, as a defense mechanism that bacteria use to protect themselves against infection. It’s a great example of a biological system with important implications for medicine, agriculture, etc. that would never have been discovered and developed as a gene modification tool without support for basic research—and it underscores the need to increase funding for basic research. Seminal discoveries have always come from unexpected places.


Please continue to progress our hurry sickness . How Many Years Will Continue this work ?
It sounds like an amazing work. Both my two sons and I have fshd. It would be amazing for this to be A great breakthrough. Wondering how this would affect someone who already has fshd. And how would it beCarried out? Thank you for all the amazing work that you’re doing!
CRISPR-based therapies would likely be developed as a form of gene therapy, in which a genetically engineered virus is designed to deliver the treatment to the DNA of the patient. The patient’s own cells would then start making the molecules that would inactivate the DUX4 gene (thought to cause FSHD). This approach requires testing to make sure the virus is safe and effective, as well as manufacturing trillions of virus to infuse into the patient.
My 11 year old son is diagnosed with FSHD and I can now see the degradation of muscles. I need help in understanding what can I do to slow down the progression and that how far away are we from developing the technology. Is there a forum where we can monitor the progress of this technology development, are we doing any clinical trials on humans?